EGRAPHS Community Call Talk#
Egglog as a tool for building an optimizing composable type safe DSLs in Python
… to help drive theoretical development of e-graphs in conjunction with impacting (large) real world communities.
Now that I have this great e-graph library in Python, what extra mechanisms do I need to make it useful in existing Python code?
This talk will go thorugh a few techniques developed and also point to how by bringing in use cases from scientific Python can help drive further theoretic research
Optimizing Scikit-learn with Numba#
We are going to work through the different pieces needed to optimize a Scikit-learn pipeline using Numba and egglog.
from __future__ import annotations
import os
import numpy as np
# Ensure SciPy array API support is enabled before importing sklearn/scipy
os.environ.setdefault("SCIPY_ARRAY_API", "1")
import sklearn
from sklearn import datasets
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
# Tell sklearn to treat arrays as following array API
sklearn.set_config(array_api_dispatch=True)
X_np, y_np = datasets.load_iris().data, datasets.load_iris().target
# Assumption: I want to optimize calling this many times on data similar to that above
def run_lda(x, y):
lda = LinearDiscriminantAnalysis(n_components=2)
return lda.fit(x, y).transform(x)
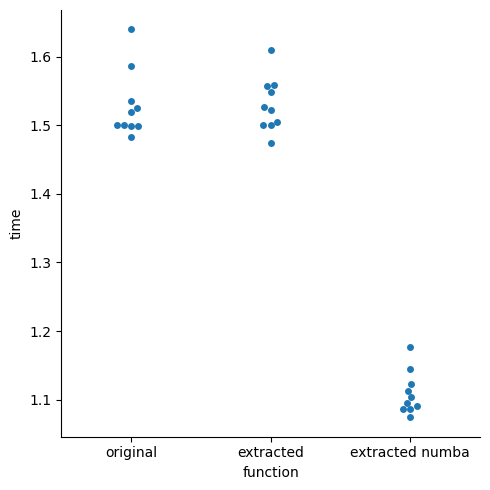
We can do this using egglog to generate Python code and Numba to JIT compile it to LLVM, resulting in a speedup:
# The first thing we need to do is create our symbolic arrays and get back our symbolic output
from egglog.exp.array_api import *
X_arr = NDArray.var("X")
assume_dtype(X_arr, X_np.dtype)
assume_shape(X_arr, X_np.shape)
assume_isfinite(X_arr)
y_arr = NDArray.var("y")
assume_dtype(y_arr, y_np.dtype)
assume_shape(y_arr, y_np.shape)
assume_value_one_of(y_arr, tuple(map(int, np.unique(y_np)))) # type: ignore[arg-type]
egraph = EGraph()
with set_array_api_egraph(egraph):
res = run_lda(X_arr, y_arr)
res
_NDArray_1 = NDArray.var("X")
assume_dtype(_NDArray_1, DType.float64)
assume_shape(_NDArray_1, TupleInt(Vec(Int(150), Int(4))))
assume_isfinite(_NDArray_1)
_NDArray_2 = NDArray.var("y")
assume_dtype(_NDArray_2, DType.int64)
assume_shape(_NDArray_2, TupleInt(Vec(Int(150))))
assume_value_one_of(_NDArray_2, TupleValue(Vec(Value.from_int(Int(0)), Value.from_int(Int(1)), Value.from_int(Int(2)))))
_NDArray_3 = astype(unique_counts_counts(_NDArray_2), asarray(_NDArray_1).dtype) / NDArray(RecursiveValue(Value.from_float(Float(150.0))))
_NDArray_4 = zeros(
TupleInt(
Vec(
NDArray(RecursiveValue.vec(Vec(RecursiveValue(Value.from_int(Int(0))), RecursiveValue(Value.from_int(Int(1))), RecursiveValue(Value.from_int(Int(2)))))).shape[Int(0)],
asarray(_NDArray_1).shape[Int(1)],
)
),
OptionalDType.some(asarray(_NDArray_1).dtype),
OptionalDevice.some(asarray(_NDArray_1).device),
)
_MultiAxisIndexKeyItem_1 = MultiAxisIndexKeyItem.slice(Slice())
_IndexKey_1 = IndexKey.multi_axis(MultiAxisIndexKey.from_vec(Vec(MultiAxisIndexKeyItem.int(Int(0)), _MultiAxisIndexKeyItem_1)))
_NDArray_5 = NDArray(RecursiveValue(Value.from_int(Int(0))))
_NDArray_4[_IndexKey_1] = mean(asarray(_NDArray_1)[IndexKey.ndarray(unique_inverse_inverse_indices(_NDArray_2) == _NDArray_5)], OptionalIntOrTuple.int(Int(0)))
_IndexKey_2 = IndexKey.multi_axis(MultiAxisIndexKey.from_vec(Vec(MultiAxisIndexKeyItem.int(Int(1)), _MultiAxisIndexKeyItem_1)))
_NDArray_4[_IndexKey_2] = mean(
asarray(_NDArray_1)[IndexKey.ndarray(unique_inverse_inverse_indices(_NDArray_2) == NDArray(RecursiveValue(Value.from_int(Int(1)))))], OptionalIntOrTuple.int(Int(0))
)
_IndexKey_3 = IndexKey.multi_axis(MultiAxisIndexKey.from_vec(Vec(MultiAxisIndexKeyItem.int(Int(2)), _MultiAxisIndexKeyItem_1)))
_NDArray_4[_IndexKey_3] = mean(
asarray(_NDArray_1)[IndexKey.ndarray(unique_inverse_inverse_indices(_NDArray_2) == NDArray(RecursiveValue(Value.from_int(Int(2)))))], OptionalIntOrTuple.int(Int(0))
)
_NDArray_6 = asarray(reshape(asarray(_NDArray_2), TupleInt(Vec(Int(-1)))))
_Int_1 = unique_values(concat(TupleNDArray(Vec(unique_values(asarray(_NDArray_6)))))).shape[Int(0)]
_NDArray_7 = concat(
TupleNDArray(
Vec(
asarray(_NDArray_1)[IndexKey.ndarray(_NDArray_6 == _NDArray_5)] - _NDArray_4[_IndexKey_1],
asarray(_NDArray_1)[IndexKey.ndarray(_NDArray_6 == NDArray(RecursiveValue(Value.from_int(Int(1)))))] - _NDArray_4[_IndexKey_2],
asarray(_NDArray_1)[IndexKey.ndarray(_NDArray_6 == NDArray(RecursiveValue(Value.from_int(Int(2)))))] - _NDArray_4[_IndexKey_3],
)
),
OptionalInt.some(Int(0)),
)
_NDArray_8 = std(_NDArray_7, OptionalIntOrTuple.int(Int(0)))
_NDArray_8[IndexKey.ndarray(std(_NDArray_7, OptionalIntOrTuple.int(Int(0))) == _NDArray_5)] = NDArray(RecursiveValue(Value.from_float(Float(1.0))))
_TupleNDArray_1 = svd_(
sqrt(
asarray(
NDArray(RecursiveValue(Value.from_float(Float(1.0) / Float.from_int(Int(150) - _Int_1)))),
OptionalDType.some(asarray(_NDArray_1).dtype),
OptionalBool.none,
OptionalDevice.some(asarray(_NDArray_1).device),
)
)
* (_NDArray_7 / _NDArray_8),
Boolean(False),
)
_Slice_1 = Slice(
OptionalInt.none, OptionalInt.some(sum(astype(_TupleNDArray_1[Int(1)] > NDArray(RecursiveValue(Value.from_float(Float(0.0001)))), DType.int32)).index(TupleInt()).to_int)
)
_NDArray_9 = (
_TupleNDArray_1[Int(2)][IndexKey.multi_axis(MultiAxisIndexKey.from_vec(Vec(MultiAxisIndexKeyItem.slice(_Slice_1), _MultiAxisIndexKeyItem_1)))] / _NDArray_8
).T / _TupleNDArray_1[Int(1)][IndexKey.slice(_Slice_1)]
_TupleNDArray_2 = svd_(
(
sqrt(NDArray(RecursiveValue(Value.from_int(Int(150)))) * _NDArray_3 * NDArray(RecursiveValue(Value.from_float(Float(1.0) / Float.from_int(_Int_1 - Int(1))))))
* (_NDArray_4 - _NDArray_3 @ _NDArray_4).T
).T
@ _NDArray_9,
Boolean(False),
)
(
(asarray(_NDArray_1) - _NDArray_3 @ _NDArray_4)
@ (
_NDArray_9
@ _TupleNDArray_2[Int(2)].T[
IndexKey.multi_axis(
MultiAxisIndexKey.from_vec(
Vec(
_MultiAxisIndexKeyItem_1,
MultiAxisIndexKeyItem.slice(
Slice(
OptionalInt.none,
OptionalInt.some(
sum(
astype(
_TupleNDArray_2[Int(1)] > NDArray(RecursiveValue(Value.from_float(Float(0.0001)))) * _TupleNDArray_2[Int(1)][IndexKey.int(Int(0))],
DType.int32,
)
)
.index(TupleInt())
.to_int
),
)
),
)
)
)
]
)
)[IndexKey.multi_axis(MultiAxisIndexKey.from_vec(Vec(_MultiAxisIndexKeyItem_1, MultiAxisIndexKeyItem.slice(Slice(OptionalInt.none, OptionalInt.some(Int(2)))))))]
In order to run this, scikit-learn treated these objects as “array like”, meaning they conformed to the Array API.
Conversions: From Python to egglog#
Use conversions if you want your egglog API to be called with existing Python objects, without manually upcasting them
We will see this in our example:
class LinearDiscriminantAnalysis:
...
def fit(self, X, y):
...
_, cnts = xp.unique_counts(y) # non-negative ints
self.priors_ = xp.astype(cnts, X.dtype) / float(y.shape[0])
Ends up resulting in this expression:
astype(unique_counts(_NDArray_3)[Int(1)], asarray(_NDArray_1).dtype) / NDArray.scalar(Value.float(Float(1000000.0)))
How?
We have exposed a global conversion logic, where if you pass an arg to egglog and it isn’t the correct type, it will try to upcast the arg to the required egglog type.
There is a graph of all conversions and it will find the shortest path from the input to the desired type and automatically upcast to that.
Example#
For example, in indexing, if we do a slice (i.e. 1:10:2), we convert this to our custom egglog Slice expressions:
class Slice(Expr):
def __init__(
self,
start: OptionalInt = OptionalInt.none,
stop: OptionalInt = OptionalInt.none,
step: OptionalInt = OptionalInt.none,
) -> None: ...
converter(
slice,
Slice,
lambda x: Slice(
convert(x.start, OptionalInt),
convert(x.stop, OptionalInt),
convert(x.step, OptionalInt),
),
)
class A(Expr):
def __init__(self) -> None: ...
def __getitem__(self, s: Slice) -> Int: ...
A()[:1:2] # Pytohn desugars this to A()[slice(None, 1, 2)]
A()[Slice(OptionalInt.none, OptionalInt.some(Int(1)), OptionalInt.some(Int(2)))]
Preserved Methods#
If you need your egglog objects to interact with Python control flow, you can use preserved methods to stop, compile, and return an eager result to Python
In the fit function in sklearn, there are complicated analysis that must be done eagerly, like this one, which
depends on knowing the priors, which are based on the counts of the classes in the training data, which we provided:
class LinearDiscriminantAnalysis:
def fit(self, x, y):
...
if xp.abs(xp.sum(self.priors_) - 1.0) > 1e-5:
warnings.warn("The priors do not sum to 1. Renormalizing", UserWarning)
self.priors_ = self.priors_ / self.priors_.sum()
That is why we have to provide the metadata about the arrays, so we can reduce this expression to a boolean, using some interval analysis:
_NDArray_1 = NDArray.var("y")
assume_dtype(_NDArray_1, DType.int64)
assume_shape(_NDArray_1, TupleInt(Int(1000000)))
assume_value_one_of(_NDArray_1, TupleValue(Value.int(Int(0))) + TupleValue(Value.int(Int(1))))
_NDArray_2 = NDArray.var("X")
assume_dtype(_NDArray_2, DType.float64)
assume_shape(_NDArray_2, TupleInt(Int(1000000)) + TupleInt(Int(20)))
assume_isfinite(_NDArray_2)
(
abs(
sum(astype(unique_counts(asarray(reshape(asarray(_NDArray_1), TupleInt(Int(-1)))))[Int(1)], asarray(_NDArray_2).dtype) / NDArray.scalar(Value.float(Float(1000000.0))))
- NDArray.scalar(Value.float(Float(1.0)))
)
> NDArray.scalar(Value.float(Float(1e-05)))
).to_value().to_bool.bool
But how does it move through the control flow if the expression is lazy? We can implement “preserved methods” which can evaluate an expression eagerly by adding it to the EGraph and evalutating it:
Example#
class Boolean(Expr):
# Can be constructed from and convert to a primitive egglog bool:
def __init__(self, b: BoolLike) -> None: ...
@property
def bool(self) -> Bool: ...
# support boolean ops
def __and__(self, other: Boolean) -> Boolean: ...
# Can be treated like a Python bool
@method(preserve=True)
def __bool__(self) -> bool:
egraph = EGraph()
egraph.register(self)
egraph.run(bool_rewrites.saturate())
return egraph.extract(self.bool).value
x = var("x", Boolean)
y = var("y", Bool)
bool_rewrites = ruleset(
rule(eq(x).to(Boolean(y))).then(set_(x.bool).to(y)),
rewrite(Boolean(True) & Boolean(True)).to(Boolean(True)),
)
expr = Boolean(True) & Boolean(True)
expr
Boolean(True) & Boolean(True)
if expr:
print("yep it's true")
yep it's true
Mutations#
Mark a function or method as mutating the first arg, to translate it to pure function, but which acts imperative.
Another pattern that comes up a lot in Python is methods that mutate their arguments. But egglog is a pure functional language, so how do we support that?
Well we can convert functions that mutate an arg into one that returns a modified value of that argument. That way, we can keep using it with existing imperative methods and things work as they should.
For example, arrays support __setitem__, and this is used by scikit-learn:
class LinearDiscriminantAnalysis:
...
def _solve_svd(self, X, y):
...
# 1) within (univariate) scaling by with classes std-dev
std = xp.std(Xc, axis=0)
# avoid division by zero in normalization
std[std == 0] = 1.0
This will be translated to the following expressions, where there will be a new array created in the graph for the modified version:
_NDArray_8 = std(_NDArray_7, _OptionalIntOrTuple_1)
_NDArray_8[ndarray_index(std(_NDArray_7, _OptionalIntOrTuple_1) == NDArray.scalar(Value.int(Int(0))))] = NDArray.scalar(Value.float(Float(1.0)))
Example#
We can see a simpler example of this below:
class ListOfInts(Expr):
def __init__(self) -> None: ...
def __getitem__(self, i: i64Like) -> Int: ...
def __setitem__(self, i: i64Like, v: Int) -> None: ...
xs = ListOfInts()
xs[0] = Int(1)
new_egraph = EGraph()
new_egraph.register(xs[0])
new_egraph.display()
Subsumption#
mark a rewrite as subsumed to replace a smaller expression with a larger one
Now that we have a program, what do we do with it? Well first, we can optimize it, running rewrites, including those to translate from Numba forms to others.
We have added “subsumption” to egglog, to support directional rewrites, so that the left hand side is not extractable and not matchable. This is handy when we want to extract a value with more expressions or a higher cost, in a particular instance:
@array_api_numba_ruleset.register
def _mean(y: NDArray, x: NDArray, i: Int):
axis = OptionalIntOrTuple.some(IntOrTuple.int(i))
res = sum(x, axis) / NDArray.scalar(Value.int(x.shape[i]))
yield rewrite(mean(x, axis, FALSE), subsume=True).to(res)
yield rewrite(mean(x, axis, TRUE), subsume=True).to(expand_dims(res, i))
We can optimize this with the numba rules and we can see this rule take place in the _NDArray_9 line:
from egglog.exp.array_api_numba import array_api_numba_schedule
egraph.register(res)
egraph.run(array_api_numba_schedule)
simplified_res = egraph.extract(res)
simplified_res
_NDArray_1 = NDArray.var("X")
assume_dtype(_NDArray_1, DType.float64)
assume_shape(_NDArray_1, TupleInt(Vec(Int(150), Int(4))))
assume_isfinite(_NDArray_1)
_NDArray_2 = NDArray.var("y")
assume_dtype(_NDArray_2, DType.int64)
assume_shape(_NDArray_2, TupleInt(Vec(Int(150))))
assume_value_one_of(_NDArray_2, TupleValue(Vec(Value.from_int(Int(0)), Value.from_int(Int(1)), Value.from_int(Int(2)))))
_NDArray_3 = astype(
NDArray(
RecursiveValue.vec(
Vec(
RecursiveValue(sum(_NDArray_2 == NDArray(RecursiveValue(Value.from_int(Int(0))))).index(TupleInt())),
RecursiveValue(sum(_NDArray_2 == NDArray(RecursiveValue(Value.from_int(Int(1))))).index(TupleInt())),
RecursiveValue(sum(_NDArray_2 == NDArray(RecursiveValue(Value.from_int(Int(2))))).index(TupleInt())),
)
)
),
DType.float64,
) / NDArray(RecursiveValue(Value.from_int(Int(150))))
_NDArray_4 = zeros(TupleInt(Vec(Int(3), Int(4))), OptionalDType.some(DType.float64), OptionalDevice.some(_NDArray_1.device))
_MultiAxisIndexKeyItem_1 = MultiAxisIndexKeyItem.slice(Slice())
_IndexKey_1 = IndexKey.multi_axis(MultiAxisIndexKey.from_vec(Vec(MultiAxisIndexKeyItem.int(Int(0)), _MultiAxisIndexKeyItem_1)))
_NDArray_5 = _NDArray_1[IndexKey.ndarray(_NDArray_2 == NDArray(RecursiveValue(Value.from_int(Int(0)))))]
_NDArray_4[_IndexKey_1] = sum(_NDArray_5, OptionalIntOrTuple.int(Int(0))) / NDArray(RecursiveValue(Value.from_int(_NDArray_5.shape[Int(0)])))
_IndexKey_2 = IndexKey.multi_axis(MultiAxisIndexKey.from_vec(Vec(MultiAxisIndexKeyItem.int(Int(1)), _MultiAxisIndexKeyItem_1)))
_NDArray_6 = _NDArray_1[IndexKey.ndarray(_NDArray_2 == NDArray(RecursiveValue(Value.from_int(Int(1)))))]
_NDArray_4[_IndexKey_2] = sum(_NDArray_6, OptionalIntOrTuple.int(Int(0))) / NDArray(RecursiveValue(Value.from_int(_NDArray_6.shape[Int(0)])))
_IndexKey_3 = IndexKey.multi_axis(MultiAxisIndexKey.from_vec(Vec(MultiAxisIndexKeyItem.int(Int(2)), _MultiAxisIndexKeyItem_1)))
_NDArray_7 = _NDArray_1[IndexKey.ndarray(_NDArray_2 == NDArray(RecursiveValue(Value.from_int(Int(2)))))]
_NDArray_4[_IndexKey_3] = sum(_NDArray_7, OptionalIntOrTuple.int(Int(0))) / NDArray(RecursiveValue(Value.from_int(_NDArray_7.shape[Int(0)])))
_NDArray_8 = concat(TupleNDArray(Vec(_NDArray_5 - _NDArray_4[_IndexKey_1], _NDArray_6 - _NDArray_4[_IndexKey_2], _NDArray_7 - _NDArray_4[_IndexKey_3])), OptionalInt.some(Int(0)))
_NDArray_9 = square(_NDArray_8 - expand_dims(sum(_NDArray_8, OptionalIntOrTuple.int(Int(0))) / NDArray(RecursiveValue(Value.from_int(_NDArray_8.shape[Int(0)])))))
_NDArray_10 = sqrt(sum(_NDArray_9, OptionalIntOrTuple.int(Int(0))) / NDArray(RecursiveValue(Value.from_int(_NDArray_9.shape[Int(0)]))))
_NDArray_11 = copy(_NDArray_10)
_NDArray_11[IndexKey.ndarray(_NDArray_10 == NDArray(RecursiveValue(Value.from_int(Int(0)))))] = NDArray(
RecursiveValue(Value.from_float(Float.rational(BigRat(BigInt.from_string("1"), BigInt.from_string("1")))))
)
_TupleNDArray_1 = svd_(
sqrt(
asarray(
NDArray(RecursiveValue(Value.from_float(Float.rational(BigRat(BigInt.from_string("1"), BigInt.from_string("147")))))),
OptionalDType.some(DType.float64),
OptionalBool.none,
OptionalDevice.some(_NDArray_1.device),
)
)
* (_NDArray_8 / _NDArray_11),
Boolean(False),
)
_Slice_1 = Slice(
OptionalInt.none, OptionalInt.some(sum(astype(_TupleNDArray_1[Int(1)] > NDArray(RecursiveValue(Value.from_float(Float(0.0001)))), DType.int32)).index(TupleInt()).to_int)
)
_NDArray_12 = (
_TupleNDArray_1[Int(2)][IndexKey.multi_axis(MultiAxisIndexKey.from_vec(Vec(MultiAxisIndexKeyItem.slice(_Slice_1), _MultiAxisIndexKeyItem_1)))] / _NDArray_11
).T / _TupleNDArray_1[Int(1)][IndexKey.slice(_Slice_1)]
_TupleNDArray_2 = svd_(
(
sqrt(
NDArray(RecursiveValue(Value.from_int(Int(150))))
* _NDArray_3
* NDArray(RecursiveValue(Value.from_float(Float.rational(BigRat(BigInt.from_string("1"), BigInt.from_string("2"))))))
)
* (_NDArray_4 - _NDArray_3 @ _NDArray_4).T
).T
@ _NDArray_12,
Boolean(False),
)
(
(_NDArray_1 - _NDArray_3 @ _NDArray_4)
@ (
_NDArray_12
@ _TupleNDArray_2[Int(2)].T[
IndexKey.multi_axis(
MultiAxisIndexKey.from_vec(
Vec(
_MultiAxisIndexKeyItem_1,
MultiAxisIndexKeyItem.slice(
Slice(
OptionalInt.none,
OptionalInt.some(
sum(
astype(
_TupleNDArray_2[Int(1)] > NDArray(RecursiveValue(Value.from_float(Float(0.0001)))) * _TupleNDArray_2[Int(1)][IndexKey.int(Int(0))],
DType.int32,
)
)
.index(TupleInt())
.to_int
),
)
),
)
)
)
]
)
)[IndexKey.multi_axis(MultiAxisIndexKey.from_vec(Vec(_MultiAxisIndexKeyItem_1, MultiAxisIndexKeyItem.slice(Slice(OptionalInt.none, OptionalInt.some(Int(2)))))))]
Program Gen#
Generate an imperative program from your e-graph with replacement rules that walk the graph in a fixed order
Now that we have a program, what do we do with it?
Well we showed how we can use eager evaluation to get a result, but what if we don’t want to do the computation in egglog, but instead export a program so we can execute that back in Python or in this case feed it to Python?
Well in this case we have designed a Program object which we can use to convert a funtional egglog expression back to imperative Python code:
from egglog.exp.array_api_program_gen import *
import numpy as np
import inspect
egraph = EGraph()
fn_program = egraph.let(
"fn_program",
EvalProgram(ndarray_function_two_program(simplified_res, NDArray.var("X"), NDArray.var("y")), {"np": np}),
)
egraph.run(array_api_program_gen_schedule)
fn = egraph.extract(fn_program.as_py_object).value
print(inspect.getsource(fn))
def __fn(X, y):
assert X.dtype == np.dtype(np.float64)
assert X.shape == (150, 4, )
assert np.all(np.isfinite(X))
assert y.dtype == np.dtype(np.int64)
assert y.shape == (150, )
assert set(np.unique(y)) == set((0, 1, 2, ))
_0 = y == np.array(0)
_1 = np.sum(_0)
_2 = y == np.array(1)
_3 = np.sum(_2)
_4 = y == np.array(2)
_5 = np.sum(_4)
_6 = np.array((_1, _3, _5, )).astype(np.dtype(np.float64))
_7 = _6 / np.array(150)
_8 = np.zeros((3, 4, ), dtype=np.dtype(np.float64))
_9 = np.sum(X[_0], axis=0)
_10 = _9 / np.array(X[_0].shape[0])
_8[0, :,] = _10
_11 = np.sum(X[_2], axis=0)
_12 = _11 / np.array(X[_2].shape[0])
_8[1, :,] = _12
_13 = np.sum(X[_4], axis=0)
_14 = _13 / np.array(X[_4].shape[0])
_8[2, :,] = _14
_15 = _7 @ _8
_16 = X - _15
_17 = np.sqrt(np.asarray(np.array(float(1 / 147)), np.dtype(np.float64)))
_18 = X[_0] - _8[0, :,]
_19 = X[_2] - _8[1, :,]
_20 = X[_4] - _8[2, :,]
_21 = np.concatenate((_18, _19, _20, ), axis=0)
_22 = np.sum(_21, axis=0)
_23 = _22 / np.array(_21.shape[0])
_24 = np.expand_dims(_23, 0)
_25 = _21 - _24
_26 = np.square(_25)
_27 = np.sum(_26, axis=0)
_28 = _27 / np.array(_26.shape[0])
_29 = np.sqrt(_28)
_30 = _29 == np.array(0)
_29[_30] = np.array(float(1))
_31 = _21 / _29
_32 = _17 * _31
_33 = np.linalg.svd(_32, full_matrices=False)
_34 = _33[1] > np.array(0.0001)
_35 = _34.astype(np.dtype(np.int32))
_36 = np.sum(_35)
_37 = _33[2][:_36, :,] / _29
_38 = _37.T / _33[1][:_36]
_39 = np.array(150) * _7
_40 = _39 * np.array(float(1 / 2))
_41 = np.sqrt(_40)
_42 = _8 - _15
_43 = _41 * _42.T
_44 = _43.T @ _38
_45 = np.linalg.svd(_44, full_matrices=False)
_46 = np.array(0.0001) * _45[1][0]
_47 = _45[1] > _46
_48 = _47.astype(np.dtype(np.int32))
_49 = np.sum(_48)
_50 = _38 @ _45[2].T[:, :_49,]
_51 = _16 @ _50
return _51[:, :2,]
From there we can complete our work, by optimizing with numba and we can call with our original values:
from numba import njit
njit(fn)(X_np, y_np)
/tmp/egglog-e3c6ab9b-1e09-4754-8cb6-65cee7e50624.py:62: NumbaPerformanceWarning: '@' is faster on contiguous arrays, called on (Array(float64, 2, 'C', False, aligned=True), Array(float64, 2, 'A', False, aligned=True))
_50 = _38 @ _45[2].T[:, :_49,]
array([[ 8.06179978e+00, -3.00420621e-01],
[ 7.12868772e+00, 7.86660426e-01],
[ 7.48982797e+00, 2.65384488e-01],
[ 6.81320057e+00, 6.70631068e-01],
[ 8.13230933e+00, -5.14462530e-01],
[ 7.70194674e+00, -1.46172097e+00],
[ 7.21261762e+00, -3.55836209e-01],
[ 7.60529355e+00, 1.16338380e-02],
[ 6.56055159e+00, 1.01516362e+00],
[ 7.34305989e+00, 9.47319209e-01],
[ 8.39738652e+00, -6.47363392e-01],
[ 7.21929685e+00, 1.09646389e-01],
[ 7.32679599e+00, 1.07298943e+00],
[ 7.57247066e+00, 8.05464137e-01],
[ 9.84984300e+00, -1.58593698e+00],
[ 9.15823890e+00, -2.73759647e+00],
[ 8.58243141e+00, -1.83448945e+00],
[ 7.78075375e+00, -5.84339407e-01],
[ 8.07835876e+00, -9.68580703e-01],
[ 8.02097451e+00, -1.14050366e+00],
[ 7.49680227e+00, 1.88377220e-01],
[ 7.58648117e+00, -1.20797032e+00],
[ 8.68104293e+00, -8.77590154e-01],
[ 6.25140358e+00, -4.39696367e-01],
[ 6.55893336e+00, 3.89222752e-01],
[ 6.77138315e+00, 9.70634453e-01],
[ 6.82308032e+00, -4.63011612e-01],
[ 7.92461638e+00, -2.09638715e-01],
[ 7.99129024e+00, -8.63787128e-02],
[ 6.82946447e+00, 5.44960851e-01],
[ 6.75895493e+00, 7.59002759e-01],
[ 7.37495254e+00, -5.65844592e-01],
[ 9.12634625e+00, -1.22443267e+00],
[ 9.46768199e+00, -1.82522635e+00],
[ 7.06201386e+00, 6.63400423e-01],
[ 7.95876243e+00, 1.64961722e-01],
[ 8.61367201e+00, -4.03253602e-01],
[ 8.33041759e+00, -2.28133530e-01],
[ 6.93412007e+00, 7.05519379e-01],
[ 7.68823131e+00, 9.22362309e-03],
[ 7.91793715e+00, -6.75121313e-01],
[ 5.66188065e+00, 1.93435524e+00],
[ 7.24101468e+00, 2.72615132e-01],
[ 6.41443556e+00, -1.24730131e+00],
[ 6.85944381e+00, -1.05165396e+00],
[ 6.76470393e+00, 5.05151855e-01],
[ 8.08189937e+00, -7.63392750e-01],
[ 7.18676904e+00, 3.60986823e-01],
[ 8.31444876e+00, -6.44953177e-01],
[ 7.67196741e+00, 1.34893840e-01],
[-1.45927545e+00, -2.85437643e-02],
[-1.79770574e+00, -4.84385502e-01],
[-2.41694888e+00, 9.27840307e-02],
[-2.26247349e+00, 1.58725251e+00],
[-2.54867836e+00, 4.72204898e-01],
[-2.42996725e+00, 9.66132066e-01],
[-2.44848456e+00, -7.95961954e-01],
[-2.22666513e-01, 1.58467318e+00],
[-1.75020123e+00, 8.21180130e-01],
[-1.95842242e+00, 3.51563753e-01],
[-1.19376031e+00, 2.63445570e+00],
[-1.85892567e+00, -3.19006544e-01],
[-1.15809388e+00, 2.64340991e+00],
[-2.66605725e+00, 6.42504540e-01],
[-3.78367218e-01, -8.66389312e-02],
[-1.20117255e+00, -8.44373592e-02],
[-2.76810246e+00, -3.21995363e-02],
[-7.76854039e-01, 1.65916185e+00],
[-3.49805433e+00, 1.68495616e+00],
[-1.09042788e+00, 1.62658350e+00],
[-3.71589615e+00, -1.04451442e+00],
[-9.97610366e-01, 4.90530602e-01],
[-3.83525931e+00, 1.40595806e+00],
[-2.25741249e+00, 1.42679423e+00],
[-1.25571326e+00, 5.46424197e-01],
[-1.43755762e+00, 1.34424979e-01],
[-2.45906137e+00, 9.35277280e-01],
[-3.51848495e+00, -1.60588866e-01],
[-2.58979871e+00, 1.74611728e-01],
[ 3.07487884e-01, 1.31887146e+00],
[-1.10669179e+00, 1.75225371e+00],
[-6.05524589e-01, 1.94298038e+00],
[-8.98703769e-01, 9.04940034e-01],
[-4.49846635e+00, 8.82749915e-01],
[-2.93397799e+00, -2.73791065e-02],
[-2.10360821e+00, -1.19156767e+00],
[-2.14258208e+00, -8.87797815e-02],
[-2.47945603e+00, 1.94073927e+00],
[-1.32552574e+00, 1.62869550e-01],
[-1.95557887e+00, 1.15434826e+00],
[-2.40157020e+00, 1.59458341e+00],
[-2.29248878e+00, 3.32860296e-01],
[-1.27227224e+00, 1.21458428e+00],
[-2.93176055e-01, 1.79871509e+00],
[-2.00598883e+00, 9.05418042e-01],
[-1.18166311e+00, 5.37570242e-01],
[-1.61615645e+00, 4.70103580e-01],
[-1.42158879e+00, 5.51244626e-01],
[ 4.75973788e-01, 7.99905482e-01],
[-1.54948259e+00, 5.93363582e-01],
[-7.83947399e+00, -2.13973345e+00],
[-5.50747997e+00, 3.58139892e-02],
[-6.29200850e+00, -4.67175777e-01],
[-5.60545633e+00, 3.40738058e-01],
[-6.85055995e+00, -8.29825394e-01],
[-7.41816784e+00, 1.73117995e-01],
[-4.67799541e+00, 4.99095015e-01],
[-6.31692685e+00, 9.68980756e-01],
[-6.32773684e+00, 1.38328993e+00],
[-6.85281335e+00, -2.71758963e+00],
[-4.44072512e+00, -1.34723692e+00],
[-5.45009572e+00, 2.07736942e-01],
[-5.66033713e+00, -8.32713617e-01],
[-5.95823722e+00, 9.40175447e-02],
[-6.75926282e+00, -1.60023206e+00],
[-5.80704331e+00, -2.01019882e+00],
[-5.06601233e+00, 2.62733839e-02],
[-6.60881882e+00, -1.75163587e+00],
[-9.17147486e+00, 7.48255067e-01],
[-4.76453569e+00, 2.15573720e+00],
[-6.27283915e+00, -1.64948141e+00],
[-5.36071189e+00, -6.46120732e-01],
[-7.58119982e+00, 9.80722934e-01],
[-4.37150279e+00, 1.21297458e-01],
[-5.72317531e+00, -1.29327553e+00],
[-5.27915920e+00, 4.24582377e-02],
[-4.08087208e+00, -1.85936572e-01],
[-4.07703640e+00, -5.23238483e-01],
[-6.51910397e+00, -2.96976389e-01],
[-4.58371942e+00, 8.56815813e-01],
[-6.22824009e+00, 7.12719638e-01],
[-5.22048773e+00, -1.46819509e+00],
[-6.80015000e+00, -5.80895175e-01],
[-3.81515972e+00, 9.42985932e-01],
[-5.10748966e+00, 2.13059000e+00],
[-6.79671631e+00, -8.63090395e-01],
[-6.52449599e+00, -2.44503527e+00],
[-4.99550279e+00, -1.87768525e-01],
[-3.93985300e+00, -6.14020389e-01],
[-5.20383090e+00, -1.14476808e+00],
[-6.65308685e+00, -1.80531976e+00],
[-5.10555946e+00, -1.99218201e+00],
[-5.50747997e+00, 3.58139892e-02],
[-6.79601924e+00, -1.46068695e+00],
[-6.84735943e+00, -2.42895067e+00],
[-5.64500346e+00, -1.67771734e+00],
[-5.17956460e+00, 3.63475041e-01],
[-4.96774090e+00, -8.21140550e-01],
[-5.88614539e+00, -2.34509051e+00],
[-4.68315426e+00, -3.32033811e-01]])
These program rewrites work by first translating the code into an intermediate IR of just program assignments and statements, and then turning this into source. It does this by walking the graph first top to bottom, recording the first parent that saw every child. Then it goes from bottom to top, building up a larger set of statements as well as an expression representing each node. If a node has already been emitted somewhere else, we record that, and we always wait for all the children to complete before moving forward. That way, we can enforce some ordering and we know that everything will be emitted only once.
Example#
Here is a small example, we might want to compile, let’s say we want a function like this:
def __fn(x, y)
x[1] = 10
z = x + x
return sum(z) + y
We can build up a functional program for this and then compile it to Python source:
from egglog.exp.program_gen import *
x = Program("x", is_identifier=True)
y = Program("y", is_identifier=True)
# To reference x, we need to first emit the statement
x_modified = Program("x").statement(x + "[1] = 10")
z = (x_modified + " + " + x_modified).assign()
res = Program("sum(") + z + ") + " + y
fn = res.function_two(x, y)
egraph = EGraph()
egraph.register(fn.compile())
egraph.run(program_gen_ruleset.saturate())
print(egraph.extract(fn.statements).value)
def __fn(x, y):
x[1] = 10
_0 = x + x
return sum(_0) + y
This happens, by first going top to bottom with the compile, which will put a total ordering on all nodes, by defining one, and only one, parent expression for each expression.
Then, from the bottom up, for each node we compute an expression string and a list of statements string.
egraph
PyObject - Python objects in EGraphs#
We can also add Python objects directly to the e-graph as primitives and run rewrite rules that call back into Python
Future Work#
The next milestone use case is to be able to optimize functional array programs and rewrite them.
To implement this we need to at least support functions as values and ideally also generic types.
Example#
There is a concrete example provided by Siu from the Numba project.
We would want users to be able to write code like this:
def linalg_norm_loopnest_egglog(X: enp.NDArray, axis: enp.TupleInt) -> enp.NDArray:
# peel off the outer shape for result array
outshape = ShapeAPI(X.shape).deselect(axis).to_tuple()
# get only the inner shape for reduction
reduce_axis = ShapeAPI(X.shape).select(axis).to_tuple()
return enp.NDArray.from_fn(
outshape,
X.dtype,
lambda k: enp.sqrt(
LoopNestAPI.from_tuple(reduce_axis)
.unwrap()
.reduce(lambda carry, i: carry + enp.real(enp.conj(x := X[i + k]) * x), init=0.0)
).to_value(),
)
Which would then be rewritten to:
def linalg_norm_array_api(X: enp.NDArray, axis: enp.TupleInt) -> enp.NDArray:
outdim = enp.range_(X.ndim).filter(lambda x: ~axis.contains(x))
outshape = convert(convert(X.shape, enp.NDArray)[outdim], enp.TupleInt)
row_axis, col_axis = axis
return enp.NDArray.from_fn(
outshape,
X.dtype,
lambda k: enp.sqrt(
enp.int_product(enp.range_(X.shape[row_axis]), enp.range_(X.shape[col_axis]))
.map_to_ndarray(lambda rc: enp.real(enp.conj(x := X[rc + k]) * x))
.sum()
).to_value(),
)
And finally could be lowered to Python as:
def linalg_norm_low_level(
X: np.ndarray[tuple, np.dtype[np.float64]], axis: tuple[int, int]
) -> np.ndarray[tuple, np.dtype[np.float64]]:
# # If X ndim>=3 and axis is a 2-tuple
assert X.ndim >= 3
assert len(axis) == 2
# Y - 2
outdim = [dim for dim in range(X.ndim) if dim not in axis]
outshape = tuple(np.asarray(X.shape)[outdim])
res = np.zeros(outshape, dtype=X.dtype)
row_axis, col_axis = axis
for k in np.ndindex(outshape):
tmp = 0.0
for row in range(X.shape[row_axis]):
for col in range(X.shape[col_axis]):
idx = (row, col, *k)
x = X[idx]
tmp += (x.conj() * x).real
res[k] = np.sqrt(tmp)
return res
Conclusion#
In this talk I have gone through some details of what is needed to connect data science users to egglog:
Overall, the idea is that if we can get egglog in more users hands, in particular for data intensive workloads where the tradeoff of time for pre-computation is worth it, than this can help drive exciting future research directions and also build meaningful useful tools for the scientific open source ecosystem in Python.
If you are building DSLs in Python, or more generally want to play with e-graphs, try out egglog-python!
Around on the e-graphs Zulip for any questions.